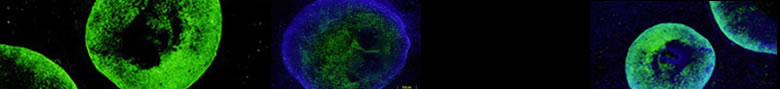
We are interested in how 10 and 30 nm chromatin fibers fold into interphase and mitotic chromosomes, how interphase chromosomes are moved and positioned within nuclei, and what this means for DNA functions such as transcription and replication. Currently, our understanding of these higher levels of chromatin organization, which we refer to as large-scale chromatin structure, is poor. We use a combination of molecular biology, cell biology, genetics, and microscopy to visualize nuclear positioning and folding dynamics of specific chromosome regions and individual gene loci and to relate this to the regulation of transcription and replication.
Most recently we have begun to apply our knowledge of chromosome dynamics to design improved tools for gene therapy and biotechnology.